Introduction
Materials and Methods
Field Surveys and Sample Collection
RNA Extraction and Multiplex RT-PCR
Sequencing and Sequence Analysis
Phylogenetic Analysis and Pairwise Comparisons
Results
Occurrence and Detection of Pear Viruses and Viroid
Occurrence of Mixed Infections
Sequence Analysis
Phylogenetic Analysis and Pairwise Comparisons
Discussion
Introduction
Belonging to the genus Pyrus, pears are commercially important fruit trees that are distributed across temperate regions in Asia, Europe, northern Africa, Australia, and New Zealand. Pear is a principal pome fruit crop in Korea, with a total cultivated area of 10,000 ha (Yoon et al., 2014; Yeam, 2016). Understanding the nature of pear pathogens is economically important. Viruses known to affect apple and pear trees include apple chlorotic leaf spot virus (ACLSV), apple stem grooving virus (ASGV), apple stem pitting virus (ASPV), and apple mosaic virus (ApMV), which are widespread and mixed infections common in many commercial cultivars of apple and pear varieties (Nemeth, 1986; Nickel and Fajardo, 2014). These latent viral infections cause significant yield reductions of up to 30%, particularly when occurring as mixed infections in Korea (Yoon et al., 2014).These viruses show a widespread distribution across apple and pear cultivation areas in various countries such as China, India, Japan, Brazil, Germany, Belarus, and Korea (Millikan and Martin, 1956; Lee et al., 2001; Pūpola et al., 2011). ACLSV, the type member of the genus Trichovirus, was first described in Malus spp. from the United States of America (Burnt et al., 1996). It is reported to have a large host range on pome and stone fruit crops, and ca have an infection rate of up to 80 - 100% in various apple cultivars (Wu et al., 1998). ACLSV induces symptoms of deformation, chlorotic leaf spots, and mosaic patterns on leaves (Katsiani et al., 2014). ASPV, a member of the genus Foveavirus, was first reported in Malus sylvestris from the Midwest in the USA and caused xylem pitting in the stem, pear vein yellowing, stony pits, and necrotic spots (Smith, 1954; Abtahi et al., 2017). Infection caused growth reductions of up to 10% in apple trees, with yield losses reaching up to 30% (Cambeli et al., 2003). ASGV, a type member of the genus Capillovirus, is associated with black necrotic leaf spot diseases in pear trees (Hong et al., 1985). ASGV is highly likely to occur especially in the ‘Niitaka’ pear cultivar and can cause serious economic damage (Nam and Kim, 1994). As expected, mixed infections by these viruses can cause substantial economic losses and induce significant yield reductions (Schmidt, 1972; Katwal et al., 2016).
Many pome fruit viruses that cause diseases are known to be transmitted by root grafts, cutting, budding, and spread through infected propagation material (Sutic et al., 1999). The production of virus-free materials and its supply to orchards is essential for effective virus control (Paunovic and Jevremovic, 2004).
Viral diseases in fruit trees present a potential danger that could damage the fruit industry and affect availability abroad. Some countries have adopted a virus protection program (VPP) to protect the spread of viral diseases (Cambeli et al., 2003; Vijeth et al., 2018). However, measures of economic losses and investigation of viral diseases in pear trees in Korea have not been reported in recent years. Hence, we determine the incidence and distribution of the most prevalent viruses infecting pear trees in the orchards as well as the molecular variability of ASSVd isolates which was the first reported viroid in pears of Korea.
Materials and Methods
Field Surveys and Sample Collection
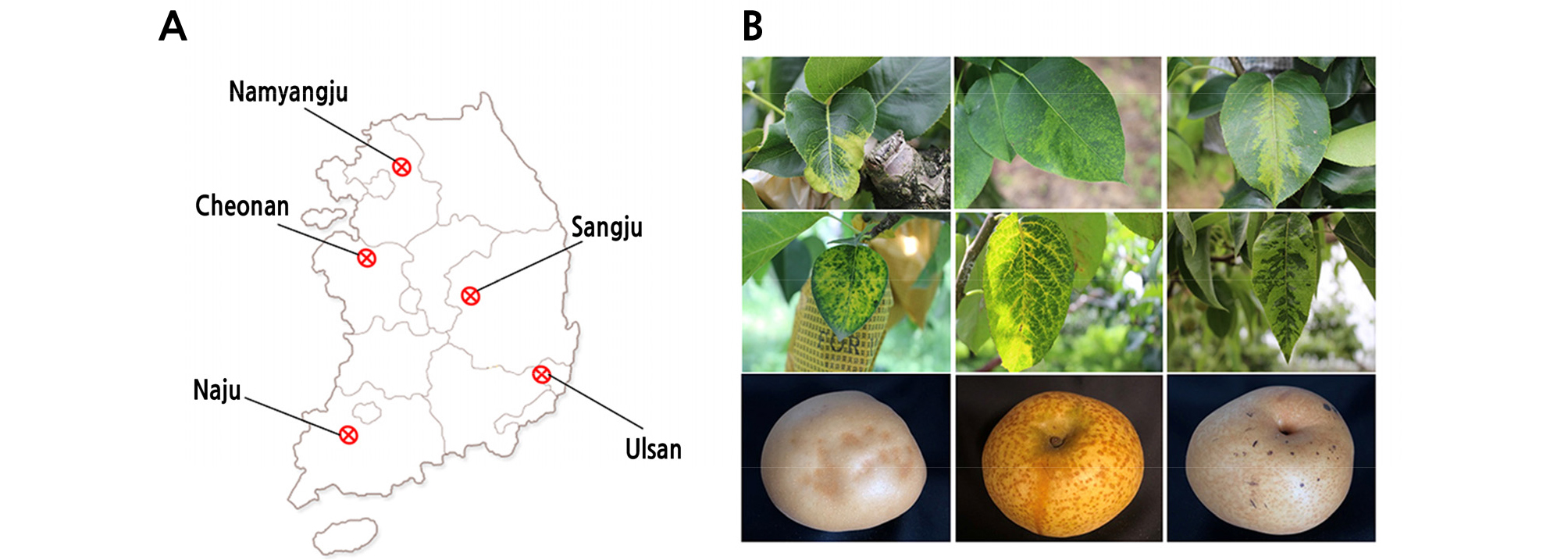
Field surveys were carried out in the spring and late summer-autumn of June 2017 to 2018. Samples of leaves and fruits were randomly collected from symptomless and symptomatic pear trees present in 56 orchards surveyed in five major pear producing areas (Sangju, Cheonan, Naju, Ulsan, and Namyangju) (Fig. 1). Leaf samples of three cultivars (Niitaka, Wonhwang, and Whangkeumbae) were taken from 363 pear trees and fruit samples of two cultivars (Niitaka and Whangkeumbae) were taken from 109 pear trees. The samples were ground in liquid nitrogen and stored at -80°C until RNA was extracted for virus detection.
RNA Extraction and Multiplex RT-PCR
Total RNA was extracted from the ground samples using the IQeasy plus plant RNA extraction Mini Kit (iNtRON, Daejeon, Korea) according to the manufacturer’s protocols. A one-tube multiplex RT-PCR was performed using a Suprime Script RT-PCR Premixture (GeNet Bio, Daejeon, Korea). Two µl of total RNA template, a forward primer, and a reverse primer for three viruses and one viroid [ACLSV, ASPV, and ASSVd (10 pmol), ASGV (2 pmol)] were added to the RT-PCR premix and filled with DEPC water up to 20 µL. Virus coat protein (CP) or partial viroid genome-specific primers used for the detection of viruses and viroid are listed in Table 1. Multiplex RT-PCR or regular RT-PCR was conducted as follows: stage 1, 50°C for 30 min; stage 2, 95°C for 5 min; stage 3, 38 cycles each of 95°C for 30 s, 58°C for 30 s and 72°C for 60 s; and stage 4, 72°C for 5 min. The PCR products were analyzed by 2% agarose gel electrophoresis in 1x TAE buffer, stained with ethidium bromide, and visualized under UV light. The size of the resultant product was determined by comparison with a 100-bp DNA ladder (iNtRON, Daejeon, Korea).
Table 1. Primer sequences and expected size for detection of pear viruses and viroid by multiplex RT-PCR and RT-PCR
Sequencing and Sequence Analysis
To confirm the identity of viruses and viroid in pear, two isolates of ACLSV, five isolates each of ASPV, and ASGV, and one isolate of ASSVd were chosen for sequencing. The RT-PCR products were purified using the MEGAquick-spinTM plus fragment DNA purification kit (iNtRON, Korea), then determined after cloning using pGEM-T Easy vector (Promega, USA), and the clones were sequenced. The sequences obtained were compared with the reference sequences for virus and viroid isolates available in GenBank database.
Phylogenetic Analysis and Pairwise Comparisons
Nucleotide sequences were aligned using CLUSTALW implemented in BioEdit. Aligned nucleotide sequences were used to construct phylogenetic trees using the MEGA 5.05 software package (Tamura et al., 2011). The phylogenetic tree was constructed based on the Kimura 2-parameter method (Kimura, 1980). Bootstrap resampling (with 1000 replications) was used to measure the reliability of individual nodes in each phylogenetic tree. Pairwise distances were calculated using the PASC algorithm available at the GenBank database (Bao et al., 2012).
Results
Occurrence and Detection of Pear Viruses and Viroid
All 363 pear leaves and 109 fruits samples collected from the five major pear producing areas (Sangju, Cheonan, Naju, Ulsan, and Namyangju) were tested using both multiplex RT-PCR and regular RT-PCR in order to decrease the error rate for virus detection. The results showed that most of the leaves carried several viral infections (Fig. 2 and Table 2). Three viruses and one viroid were detected in the leaf and fruit samples collected from all five locations. However, ApMV was not detected in pear samples. Across the regions studied, the highest infection rate was seen for ASGV (95.3%), followed by ASPV (31.7%), and ACLSV (13.8%). Meanwhile, the distribution of ASSVd (3.6%) was minor. Three viruses and one viroid were detected in the ‘Niitaka’ cultivar samples collected from all five locations, while none of the samples from ‘Whangkeumbae’ were positive for ACLSV, ASPV, and ASSVd, and none of the samples of ‘Wonhwang’ cultivar were positive for ASSVd.
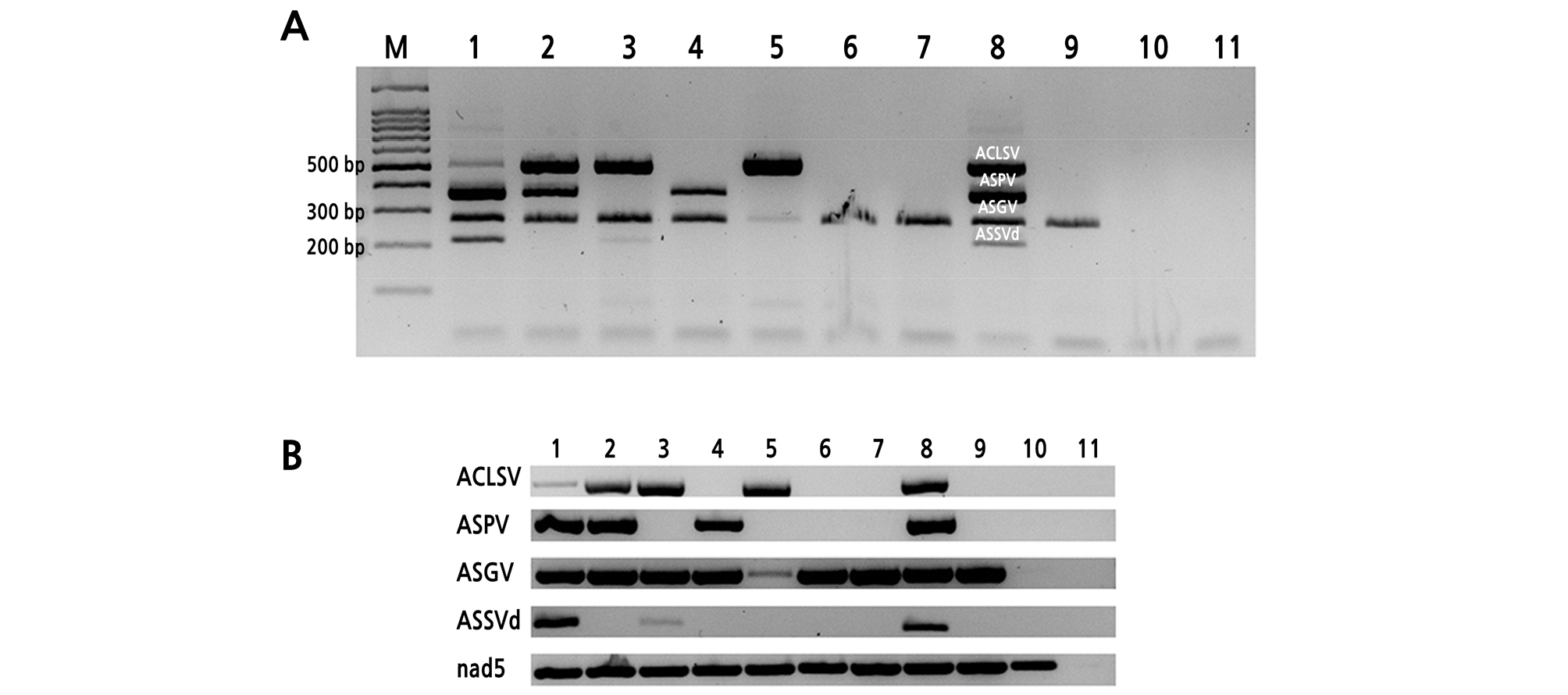
Fig. 2.
Agarose gel electrophoretic analysis of one-tube multiplex RT-PCR products (A) and regular RT-PCR products (B) amplified from total RNA preparations of virus-infected pear leaf tissues. RT-PCR products amplified from samples infected with different pome fruit viruses and viroid (lines 1-9); non-infected pear leaf (line 10); and no template control (line 11). The bands corresponding to each virus have the following lengths: 202 bp for apple scar skin viroid; 273 bp for apple stem grooving virus; 367 bp for apple stem pitting virus; 509 bp for apple chlorotic leaf spot virus. The nad5 gene was used as an internal control. The size of the nad5 gene is 181 bp.
Table 2. The incidence of viral infection in pear leaves surveyed from different locations and pear cultivars by multiplex RT-PCR
yThe conformity of positive and negative results obtained by multiplex RT-PCR, calculated in percentages.
Three viruses and one viroid were detected in the pear fruit samples (Table 3). Across the different regions, the highest infection rate was seen for ASGV (99.0%), followed by ASPV (28.4%), and ASSVd (13.8%); the distribution of ACLSV (2.8%) was minor. ACLSV was detected only in Namyangju location, while ASPV, ASGV, and ASSVd were detected in all five locations.
Table 3. The incidence of viral infection in pear fruits surveyed from different locations and pear cultivars by multiplex RT-PCR
yThe conformity of positive and negative results obtained by multiplex RT-PCR, calculated in percentages.
Occurrence of Mixed Infections
Mixed infections were found in 133 leaf samples (36.6%) and 43 fruit samples (39.4%). In leaves, 99 samples were doubly infected, 31 samples were triply infected, and a quadruple infection was observed on three leaf samples. Seven different combinations of mixed virus infections were detected (Table 4). The highest incidence (approximately 22.0%) was recorded for the double infections of ASPV and ASGV, followed by triple infection of ACLSV, ASPV, and ASGV (7.2%), and finally for double infections of ACLSV and ASGV (3.9%). In fruit, 40 samples were doubly infected and three samples were triply infected. Five different combinations of mixed virus infection were detected (Table 5). The highest incidence (23.9%) was recorded for the double infection of ASPV and ASGV, followed by double infections of ASGV and ASSVd (11.0%) and ACLSV and ASGV (1.8%).
Table 4. Single and mixed viral infection in pear leaves surveyed from different locations and pear cultivars by multiplex RT-PCR
yThe conformity of positive and negative results obtained by multiplex RT-PCR, calculated in percentages.
Table 5. Single and mixed viral infection in pear fruits surveyed from different locations and pear cultivars by multiplex RT-PCR
Sequence Analysis
The amplified RT-PCR products of the partial or complete coat protein (CP) coding sequences were determined after cloning and were sequenced to confirm the identity of the viruses and viroid detected in pear. Two CP sequences of ACLSV isolates were obtained (LC367336 and LC367337) and were most closely related (98% identity) to an ACLSV sequence (KC404868) from China. Five CP sequences of ASPV isolates were obtained (LC 367338, LC367339, LC367340, LC367341, and LC367342) and were most closely related (96% identity) to an ASPV sequence (JX673819) from China. Five CP sequences of ASGV isolates were obtained (MG382506, MG682507, MG682508, MG682509, and MG682510) and were most closely related (99% identity) to an ASGV sequence (JN792471) from South Korea. One complete ASSVd sequence was obtained (MG602681) and was most closely related (99% identity) to an ASSVd sequence (KP772686) from China. The complete genome of Korea ASSVd isolate is 331 nucleotides in length.
Phylogenetic Analysis and Pairwise Comparisons
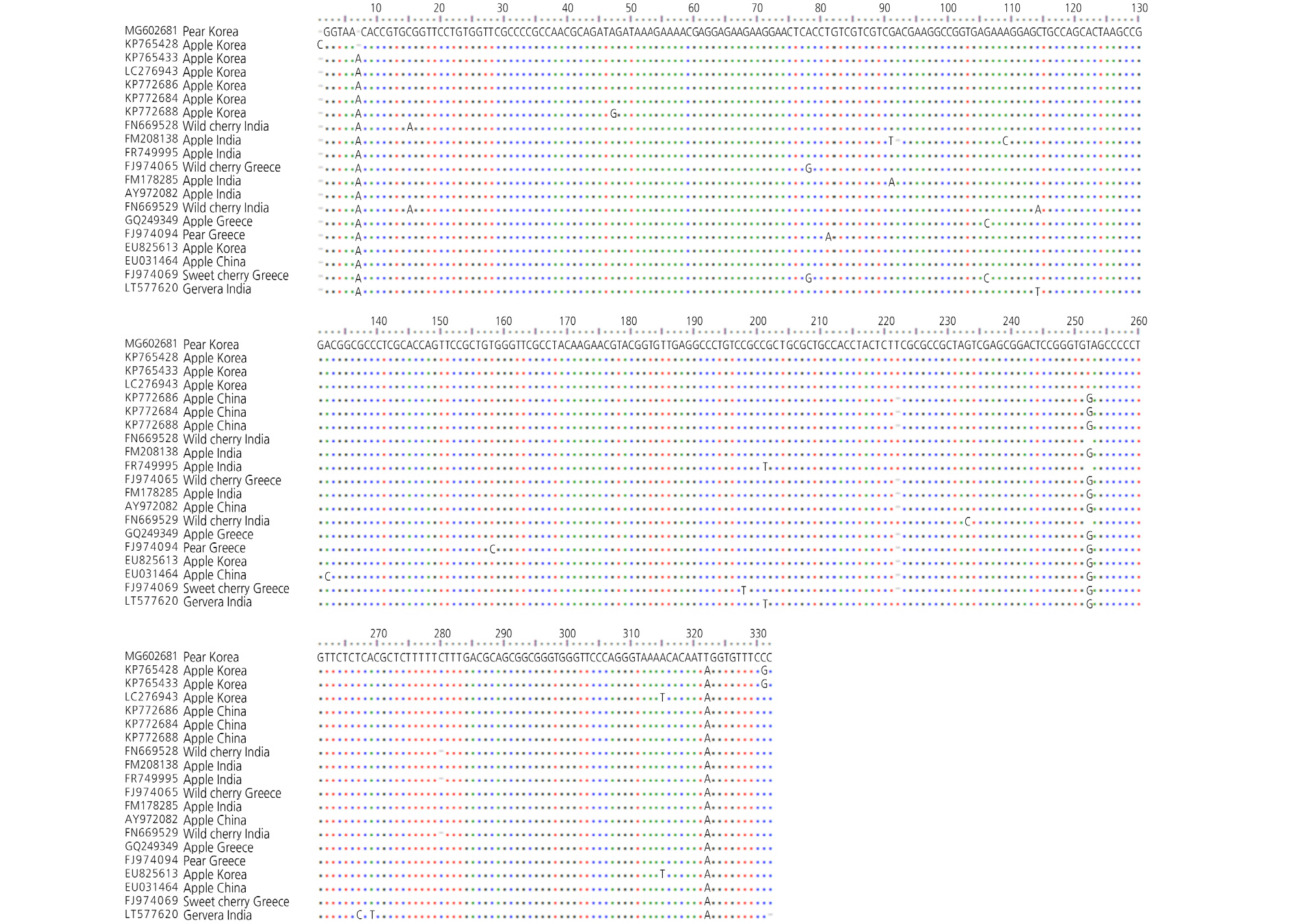
To compare the nucleotide sequence identity, a complete sequence of Korea ASSVd isolate from pear was aligned using CLUSTALW implemented in BioEdit with the complete nucleotide sequence identities of the genome of ASSVd with 19 worldwide isolates (LC276943, EU825613, FJ974094, FM208138, FM178285, FJ974065, FJ970469, GQ249349, KP772686, KP772688, AY972082, KP772684, EU031464, KP765428, KP5433, FN669528, FN669529, FR749995, and LT577620) (Table 6). The full genome of the isolate was highly conserved (Fig. 3). Pairwise distances were calculated using the PASC algorithm available at the GenBank database (Bao et al., 2012). The complete nucleotide sequence identity between isolates ranged from 98% to 99% (ASSVd) (Fig. 4). To determine the genetic relationships among the 20 isolates, we aligned nucleotide sequences and constructed a phylogenetic tree using the complete ASSVd sequences. It turns out that the Korea ASSVd isolate was placed in a different clade than the other isolates (Fig. 5). Phylogenetic analysis of the complete sequence of ASSVd and other members of genus Apscaviroid showed that the ASSVd isolate that was detected in pears is more closely related to other ASSVd isolates than to members of other viroid species. To further determine the recombination event with 20 other isolates from worldwide for genetic variation, we conducted recombination detection analysis using RDP4 software package (Martin et al., 2005). However, there were no apparent recombination events in Korea ASSVd isolate (data not shown).
Table 6. Information of apple scar skin viroid (ASSVd) isolates with complete sequences that were used in this study
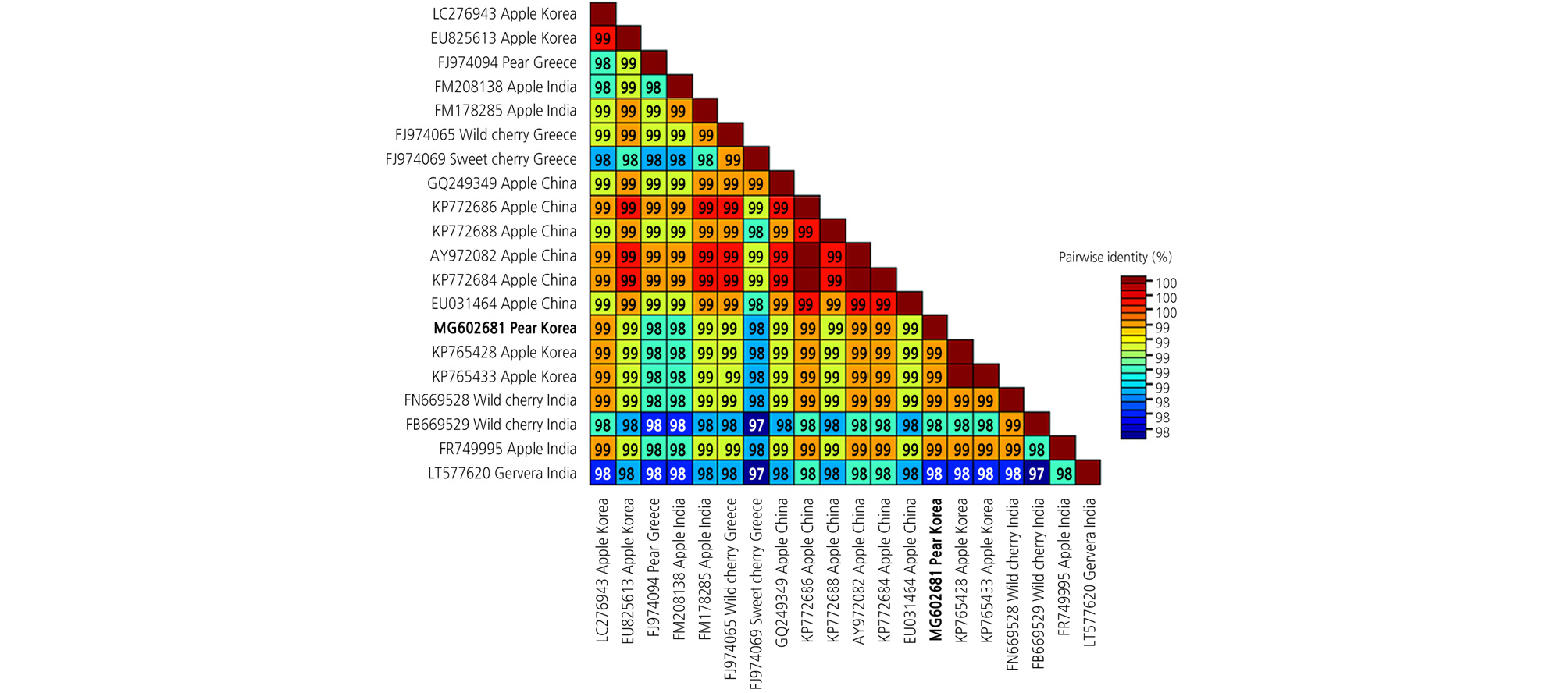
Fig. 4.
Pairwise comparisons of the complete nucleotide sequence of the apple scar skin viroid (ASSVd) genome. Information on the accession number, geographic location, and host plant for each ASSVd isolate is listed in order of the taxon names. Each number represents pairwise identities between the corresponding isolates.
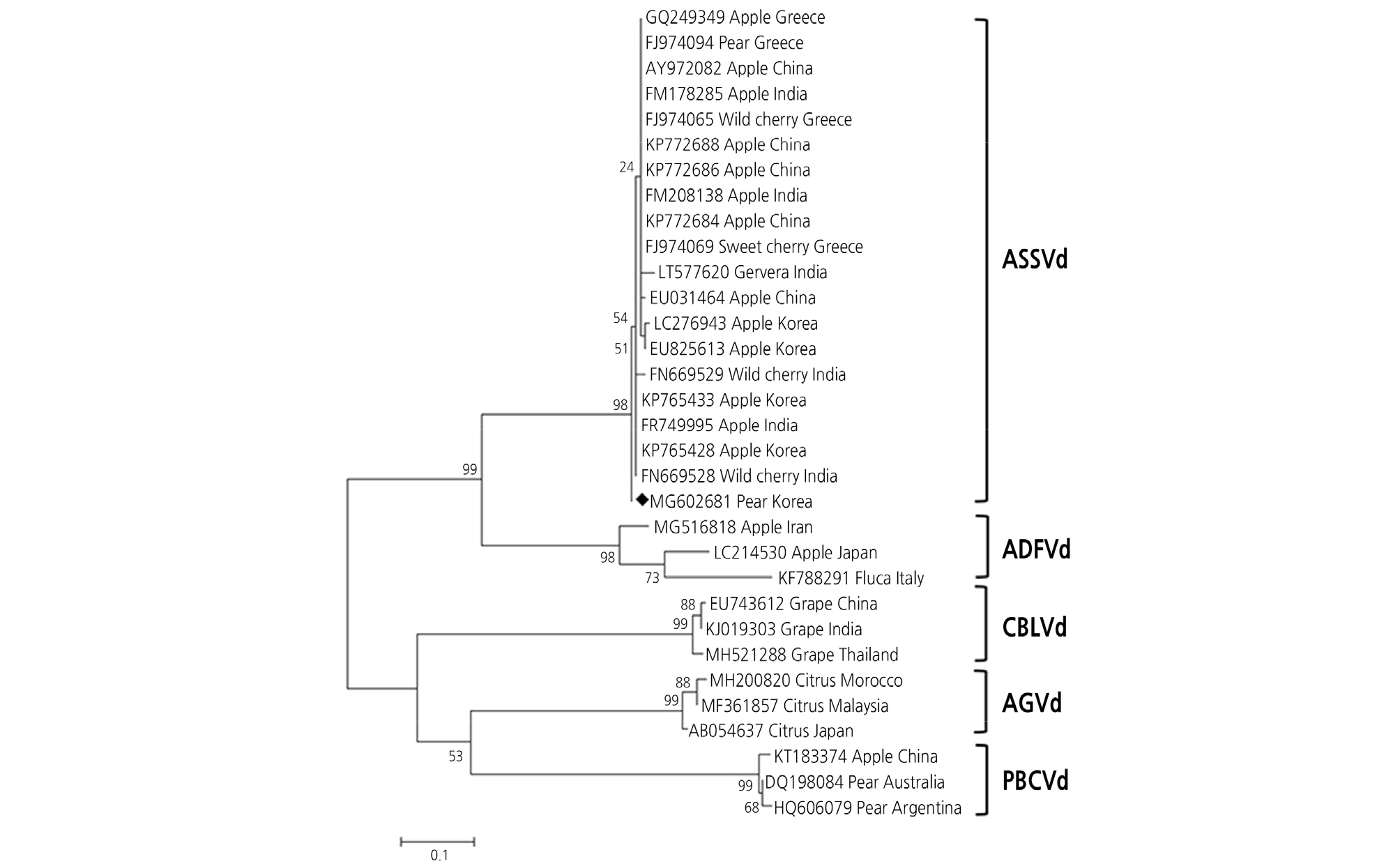

Fig. 5.
Phylogenetic tree of the 32 Apscaviroid nucleotide acid sequences of the complete gene inferred from MEGA 5.05. Each sequence is labelled with the GenBank accession number. Bootstrap test with 1,000 replicates was performed to evaluate the significance of the internal branches. Apple dimple fruit viroid : ADFVd; citrus bent leaf viroid : CBLVd; Australian grapevineviroid : AGVd; pear blister canker viroid : PBCVd.
Discussion
This study investigated the occurrence and distribution of viruses and viroid in pears in five major producing areas of Korea. These areas were surveyed in 2017 and 2018 and symptoms of chlorotic leaf spots, mosaic patterning, vein yellowing, and black necrotic leaf spots were observed. These symptoms were similar to those previously reported from virus-infected apple and pear plants in other countries (Mahfoudhi et al., 2013; Abtahi et al., 2017; Megrelishvili et al., 2017). The viruses ACLSV, ASPV, ApMV, and ASGV in apple and pear orchards are widespread (Pūpola et al., 2011). The occurrence of these viruses reported in other studies varied from 16 to 89% for ACSLV, 24 to 100% for ASPV, and 25 to 85% for ASGV (Pūpola et al., 2011; Keshavarz and Shams-Bakhsh, 2014; Hu et al., 2017; Skhiri et al., 2017). Our results indicate that of the three viruses and one viroid detected from pear crop grown in the commercial orchards in Korea (Tables 2 and 3), the infection rate of ASGV (89 to 100%) was higher than that of ASPV (14 to 45%) and ACLSV (8 to 30%). Unexpectedly, ApMV was not detected in this study. Similar results have been reported in previous studies, where ASGV was the most prevalent virus when compared to other viruses (Cho et al., 2010). We also investigated the ASSVd infection rate in various pear cultivars in Korea. ASSVd, which is a member of genus Apscaviroid, family Pospiviroidae, was first reported from China in the 1930s and also occurs in other countries such as Japan, India, Korea, and regions of Europe and North America (Ohtsuka, 1938; Millikan and Martin, 1956; Cambell et al., 1976; Lee et al., 2001; Kwon et al., 2002). Scar skin and dapple symptoms were observed on apples and rusty skin disease and fruit crinkle disease symptoms were observed on pears following infection by ASSVd (Kyriakopoulou et al., 2001; Koganezawa et al., 2003). Our data showed that ASSVd was detected only in the ‘Niitaka’ cultivar and the infection rate ASSVd in pear fruit (13.8%) was higher than that in pear leaves (1.2 to 4.9%). These infection rates of ASSVd are higher than the infection rate (1.7%) observed in 2010 (Kim et al., 2010).
Mixed infection of the three viruses (ACLSV, ASPV, and ASGV) is not a new finding. Similar mixed infections were observed in pears in 2009 in Korea (Cho et al., 2010). However, mixed infection of the three viruses and one viroid (ACLSV, ASPV, ASGV, and ASSVd) observed in this study is the first reported case in pear trees. The highest mixed infection rate of the two viruses (ASPV and ASGV) was observed in pear leaves and fruit, whereas the most common co-infection in apple was ACLSV with ASPV (Pūpola et al., 2011). Our results indicate that ASPV and ASGV are prevalent viruses in pears.
Molecular characterizations and genetic diversity of pome fruit viruses have been reported recently (Dhir et al., 2013; Jo et al., 2018). Studies in Belarus showed high genetic diversity of isolated ACLSV genomes with 84.2% nucleotide similarity to fragments sequenced from apple varieties (Kuzmitskaya et al., 2016). ASPV has also shown strong variation in the CP gene sequences in Korea (Yoon et al., 2014). Phylogenetic relationships using the genomic sequences of different ASGV isolates collected from around the world showed that 54 ASGV isolates were separated into two groups and no host grouping was seen in their phylogenetic tree (Liu et al., 2013). Our results confirmed the identity of three viruses and one viroid, which were detected in pears. These virus CP sequences and the complete sequence of viroid showed 96 - 99% identity with other viral isolates. In particular, ASSVd is more closely related to ASSVd isolates than to members of other viroid species in the phylogenetic analysis of complete sequences. In addition, results of pairwise comparison of the complete sequence of ASSVd indicated that this sequence is more closely related to ASSVd isolates (99%) than to other members of the genus Apscaviroid. The phylogenetic study of ASSVd isolates worldwide based on complete sequences showed that ASSVd isolates clearly separated from all others with a high bootstrap value. This indicated that the ASSVd clusters have no correlation to host or geographical origins. Although viruses infecting pear tree show various recombination events in their genome, ASSVd did not show recombination events, indicating that the ASSVd genome is highly conserved in the different isolates (Liu et al., 2013).
For control of fruit virus diseases, several different strategies have been developed including detection and eradication of virus-infected plants, grafting technology, eradication programs, and transgenic methods (Rowhani et al., 1995; Mathews, 2010). Control methods depend on proper identification of diseases and the causal agents. Therefore, early diagnosis and monitoring of crops are among the most important means to control temperate fruit plant viruses and viroids (Kovalskaya and Hammond, 2014; Loebenstein and Katis, 2014). Our findings indicate that three viruses and one viroid are gradually spread in Korea pear cultivation fields, which should help to minimize the possible spread of potentially harmful viruses and viroids.
Viruses and viroids of pome fruit spread mainly through vegetative propagation and grafting. ACLSV, ASPV, ASGV, and ApMV do not have any known insect vectors and are not transmitted through seeds (Pedrazzoli et al., 2008). These viruses are transmitted when a virus-infected scion bud is grafted onto a healthy rootstock or onto a previously virus-infected rootstock. Thus, occurrence of viral distribution among pear cultivars might be related to their transmission pathways. It is important to continue investigations and analyses of the occurrence of viral infections, as such studies may contribute the success of certification programs for pome fruits in Korea.